Rift Valley fever (RVF) is a mosquito borne, viral zoonosis that causes significant economic and public health impacts in ruminants and humans. Kenya has experienced several outbreaks notably in (1997–1998) and (2006–2007). RVF is caused by a Phlebovirus transmitted primarily by aedes sp mosquitoes. it is ranked among the top five priority zoonotic diseases (PZD) in Kenya based on ability to cause epidemics, economic losses and severity of infection in humans. Since May 2018, Kenya has experienced multiple RVF outbreaks in high risk counties with unexpected outbreaks in low endemic areas of western Kenya. A small-scale human-animal integrated One Health Surveillance System is proposed to enhance national decision support tools that will lead to faster detection, effective response, prevention and targeted control of RVF during the interepidemic period.
Objective
- i) to determine the seroprevalence of RVF infection in (cattle, sheep and goats) and the seroprevalence in humans and risk factors for seropositivity to RVFV ii) to characterize the distribution and diversity of potential RVF vectors and evaluate the evolutionary diversity of RVFV strains circulating in Kenya iii) To develop spatial predictive models that will contribute to the existing national risk maps for RVF.
Methodology
Humans
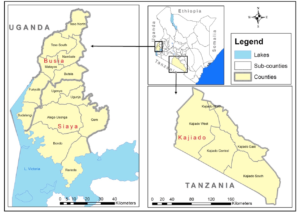
Using a hospital-based prospective and multicenter cross-sectional survey, febrile patients attending selected health facilities will be targeted, their clinical history will be obtained and physical examination performed on only consenting patients. Blood samples will be collected by a phlebotomist and serum samples tested for exposure to anti-RVF immunoglobulin G IgG/M (IgM). Busia and Siaya County referral hospitals will be selected while the Loitoktok subcounty hospital will be selected in Kajiado County. (Cases = Controls = )Livestock
Cluster sampling: The primary sampling unit for livestock will be a village (aggregation of several homes). Five villages will be selected from each of the three counties via two stage cluster sampling technique, with counties selected a priori and subcounty and village subsequently selected randomly, based on livestock population proportional to size. Cross-sectional sampling of livestock will be done, the sample size will be determined based on existing sampling methods. A total of 1200 samples will be collected among the three species (Cattle -400, Sheep -400, Goats -400)
Entomological surveillance
Mosquitos will be collected from areas where confirmed or suspected RVF cases were recently reported. Outdoor sampling will be done using CO2 -baited CDC light traps set in the evening between 18:00 to 06:00hrs the next morning. All collection sites will be mapped by a handheld GPS.
Serological and molecular analysis
The livestock antibodies will be detected using a multispecies, commercial indirect competitive ELISA (cELISA) (ID Screen, IDVet). Sera from febrile patients will also be tested by indirect ELISA for presence of anti-RVFV IgG antibodies according to established methods. Samples that will be IgM positive will be further tested by RT-PCR.
Phylogeography
Using Next-generation sequencing (NGS), a complete genome of RVFV strains in the three locations will be analyzed using established methods. Libraries for NGS will be prepared using the NEB- Next Ultra RNA kit and sequenced on an Illumina MiSeq cycle kits. Genomes will be assembled using Geneious v9.1.2 and viral-ngs with a custom RVFV database.Genomic sequences of RVFV strains/segments will be in deposited in GenBank under specific accession numbers. A maximum likelihood phylogenetic tree will be constructed for human and animal isolates for each genome segment an compared to strains from earlier outbreaks in Kenya, Uganda and Somalia
Virus Isolation from mosquitoes
Mosquitoes will be identified used standard taxonomic keys and virus isolation done by inoculating 100 μL of each blood suspension onto confluent Vero-E6 monolayers and incubated at 37°C in 2% in minimum essential medium (FBS MEM) for 14 days. This will be monitored daily for cytopathic effects (CPE) by immunofluorescence.
Expected Outputs
- Serological evidence of RVF in humans and animals in low endemic areas may demonstrate the subclinical circulation of RVFV.
- The complete genome sequences of the RVFV human and livestock strains will contribute to the understanding of the dynamics of RVFV evolution and adaptation in Kenya
- RVF risk maps and models will predict the potential areas of elevation and transmission of RVFV in Kenya
One Health impact
This investigation of Host -Vector-Pathogen interactions will have immediate impact by informing the transmission dynamics of RVF in low endemic areas and hence provide a one-health framework to be emulated by other researchers for surveillance of other endemic viral zoonoses. This multidisciplinary approach will strengthen research capacity and harness the synergy in genomic and spatial epidemiology, public health, entomology and social sciences. The tools (risk maps) may be adopted in national policies and one health stakeholders (FAO,OIE,WHO for effective preparedness and response.
